
On BIOQIC Day, PhD students gave their first presentations about their projects, the day was of informal nature.
- For each student 20 minutes was scheduled: 10 minutes for the presentation (PowerPoint) and 10 minutes for the discussion
- The PhD committees for each Student evaluated the project progress
- The sessions were open to all interested (international) scientists, which could help to fine-tune the projects
Informal meetings of the PhD students with the PhD committees took place during the breaks.
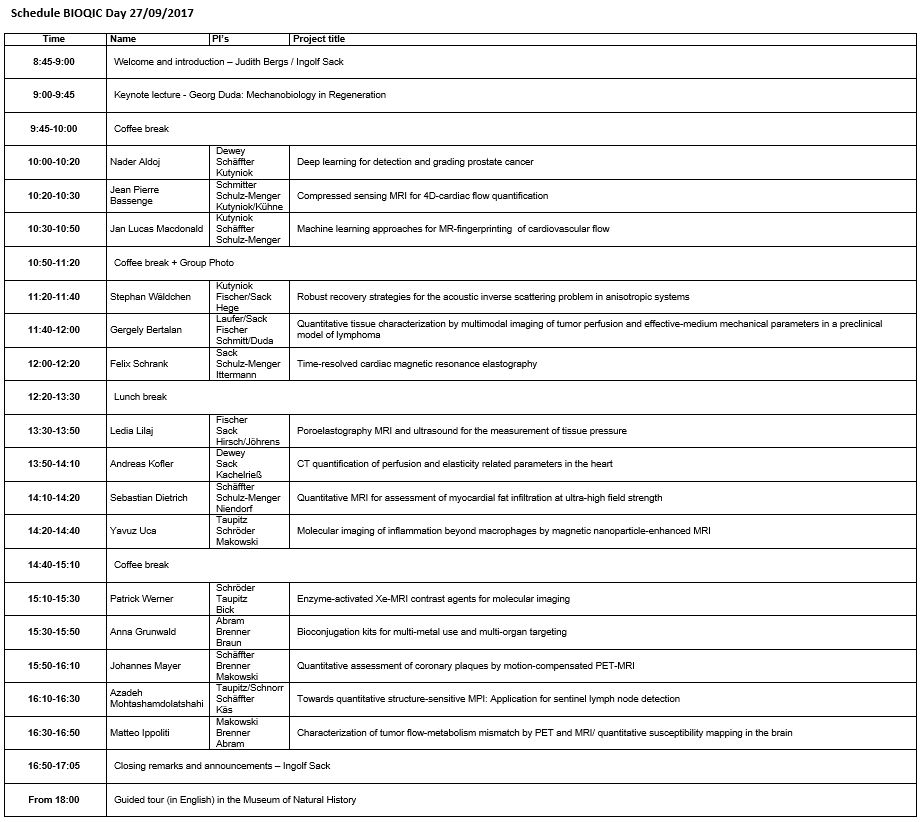
Click here for a printable PDF version of the schedule and here to download the map of the event.

BIOQIC Day 2017 was followed by the first MRE workshop from September 28th-29th in Berlin (please note: the ISMRM does not accept or assume any liability for this workshop as this workshop is not supported or organized by the ISMRM). For more information, see our website.
Many thanks to all of those who contributed to a successful BIOQIC Day!
Student abstracts BIOQIC Day 2017
Nader Aldoj
Deep learning of MRI for detection and grading of prostate cancer
PIs: Dewey, Schäffter, Kutyniok
Application Area: Cancer
Modality: MR
Background: The most common cancer type among Male population worldwide is prostate cancer. Prostate cancer can be diagnosed and graded with many different ways, e.g. Gleason score or MRI/US images, where each of those methods has its own advantages and disadvantages. For instance, taking a biopsy and using Gleason score to assess the severity and significance comes along with many complications because taking a biopsy is an invasive and painful procedure and has many complications, not to mention that the pathology assessment process in all methods is very subjective and prone to human error and misdiagnoses.
Aim: The essential aim of this project is to provide a better way of prostate cancer assessment that gets rid of the biopsy complications, the subjective assessment and replace it with an automatic- non-invasive method for localization and grading of the lesion from MRI images.
Methods: For the last years, deep learning has become a famous and promising field in recognition and classification of natural objects. For instance, convolution neural networks has the power to build up hierarchically very complex features of the object(s) being tackled and fits a good classification model that sometimes is able to classify things that human beings might miss. The idea is to use such a complex network on cancer/non-cancer in addition to clinically significant/non-significant lesions in prostate MRI images so that the network can fit a hierarchical model which has the ability to extract high level features of the lesions automatically and then assign a class depending on the annotation maps which are provided beforehand in the training step. Once the model reaches good fitting parameters, then it is able to predict the lesion localization and grading with a certain value of accuracy which in turn depends on many parameters, e.g. the number of training images, the heterogeneity of the lesion, the accuracy of the annotation maps and many more. This Model, once achieved, saves time, efforts and avoids the subjectivity of human examiners.
Jean Pierre Bassenge
Compressed sensing MRI for 4D-cardiac flow quantification
PIs: Jeanette Schulz-Menger, Sebastian Schmitter, Gitta Kutyniok
Application area: MRI
Modality: Cardiovascular
Background: 4D-flow MRI is of growing interest in different fields of cardiovascular medicine at all stages of life, as it provides new insights into hemodynamics, e.g. in the ascending aorta in subjects with bicuspid aortic valve disease or in patients with outflow tract stenosis. Furthermore, quantification of vessel wall injury by flow measurements may give insights into myocardial injury, for example in vascular disease. Nevertheless, 4D-flow MRI still suffers substantially from limited temporal and spatial resolution, and from long image acquisition times.
Aim: The objective of this PhD project is to increase the temporal and spatial resolution of 4D-flow MRI while limiting the acquisition time. This will be achieved by compressed sensing (CS) techniques, which exploit the inherent sparsity of MRI signals in different transformation domains. Different k-space sampling schemes as well as different CS reconstruction techniques such as shearlets based or dictionary learning techniques will be implemented and their performance will be quantitatively assessed. The accuracy and reproducibility of the quantitative values will be evaluated in experiments with pulsatile flow phantoms, in comparison to standard 4D flow imaging techniques. A CS based imaging protocol will then be applied in-vivo in a study with healthy volunteers, followed by a clinical pilot study including patients suffering from cardiovascular diseases.
Methods: Whereas the in-vivo studies are envisaged to be conducted primarily on a 3 Tesla MRI system, an additional focus will be set on investigating the advantages of higher magnetic field strengths (7 Tesla) for the developed CS methods. It is expected that the higher SNR and the stronger spatial variations of the receive sensitivity profiles will benefit the reconstruction methods and therefore will enhance the accuracy.
Gergely Bertalan
Quantitative tissue characterization by effective-medium mechanical parameters in preclinical small animal models
PIs: Sack, Fischer, Schmitt/Duda
Application Area: Cancer
Modalities: MRI, MRE
Background: Mechanical parameters of soft tissues sensitively change due to many pathological processes such as stroke, neurodegenerative disorders, tumors or fibrosis. In recent years, attempts to study the mechanical properties of soft tissues, through small animal models by nanoindentation methods and in vivo elastography, have begun to increase our understanding of the biophysical mechanisms underlying the development and disease of soft tissues. However, despite the significant progress achieved over the last two decades, recent modalities for in-vivo mechanical tests in small animal models remain limited in terms of spatial resolution and reproducibility.
Aim: In this project we aim to develop methods for quantitative 3D imaging including elasticity mapping in small animal models.
Methods: Preclinical MRI elastography (MRE) will be developed towards multifrequency high-resolution MRE in the mouse. Heterogeneity of mechanical parameters in neuronal tissue and tumors will be studied in animal models including a lymphoma mouse model. MRE will be correlated with histological markers of cellular and ECM components in order to conclude from macroscopic mechanical parameters to specific microstructural changes associated with disease.
Sebastian Dietrich
Quantitative MRI for assessment of myocardial fat infiltration at ultra-high field strength
PIs: Tobias Schäffter, Jeanette Schulz-Menger, Thoralf Niendorf
Application Area: Cardiovascular
Modality: MRI
Background: The concept of fatty myocardium has received attention because of its role in cardiomyopathy and obesity. Furthermore, a relationship between severity of atrial fibrillation and epicardial fat and myocardial fat infiltration has recently been reported. The healthy heart obtains approximately 70% of its energy from oxidation of long-chain fatty acids (FA). Under normal conditions, FAs are sequestered as triglycerides (TGs) and stored in adipocytes as lipid droplets, with a small amount also stored in non-adipose tissues, such as the myocardium. There is increasing evidence suggesting altered myocardial substrate utilization and subsequent excessive TG accumulation (steatosis) in myocardial disease. The concept of fatty myocardium has received attention because of its role in cardiomyopathy.
Aim: We suggest that ultra-high-field MRI allows quantification of fat infiltration in correlation with functional changes in cardiomyopathy.
Methods: During this project, a novel high-resolution 3D chemical shift imaging (CSI) sequence will be developed that enables acquisition of water and fat images of at least two cardiac phases as well as TG quantification within the myocardium at 7 Tesla. Advanced, multi-slice B0 and B1+field mapping methods will be modified to deal with respiratory and cardiac motion-induced field changes. Parallel transmission (pTX) methods will be included to counteract heterogeneous flip angle distributions at 7 Tesla. Furthermore, advanced reconstruction algorithms will be developed for multi-frequency fat spectrum modelling. A series of N images taken with small echo times (TE) increments ΔTE is the basis for CS artifact removal. The spectral bandwidth b of this acquisition is given by b = 1/ΔTE, while the spectral resolution is determined by Δf = 1/(N ΔTE). For a known and fixed MR spectrum, the spectral resolution and bandwidth can be adjusted such that each resonance line coincides with one of the N spectral windows. If the chemical shift δ of a component exceeds the bandwidth, it is aliasing back into the encoded spectral region. In an ideal case, the number of necessary echo-time increments equals the number of resonance lines. With the acquired data, we will perform a voxel-wise FT in the TE direction. All CS components will be separated and due to the known high resolution 7T spectrum and the pixel bandwidth, the CS can be compensated by sub-pixel translations. With the proposed approach, we will perform efficient imaging of TG. To validate the result we will built a dedicated phantom containing multiple oil-filled vials of different sizes. The phantom will allow for fat quantification of tiny structures that mimic tiny fat infiltration in tissue. Finally, a small clinical feasibility study is planned.
Anna Grunwald
Bioconjugation kits for multi-metal use and multi-organ targeting
PIs: Abram, Brenner, Buchert
Application Area: Cancer, cardiovascular
Modality: SPECT, PET
Background: Classical radiopharmaceuticals have usually been developed and optimized for one specific nuclide and one target organ. This was motivated by the dominance of only a few radioactive isotopes – the ‘workhorses’ – in such applications as well as the fact that the major diagnostic targets (myocardium, brain, hepatobiliary and renal tracts) ensure high numbers of annual administrations. In recent years, however, hybrid imaging tools and bioconjugation approaches in tracer chemistry have generated a need for modular tracers based on multiple radioactive nuclides. This concept allows the combination of different tracer-specific imaging techniques. Furthermore, modular tracer chemistry could support theranostic approaches, which require the quantification of therapeutic isotopes or non-radioactive drugs by medical imaging techniques. Chelator systems based on various donor atom constellations have been proven flexible and robust for stabilizing different radiometals in different chemical environments tailored for different emission tomography modalities such as PET and SPECT.
Aim: The aim of this project is to develop a novel ligand system which can coordinate various radiometals for PET and SPECT imaging as well as various target-seeking biomolecules like peptides, for instance.
Methods: Tri- to hexadentate ligands are scheduled to be synthesized on the basis of thiocarbamoylbenzimide chlorides, aroylthioureas or aminoalcohols. Each of the ligands will be synthesized in a way that allows orthogonal bioconjugation. For this, carboxylic or alkyne residues are attached in appropriate positions so that biomolecules are bound via peptide coupling or ‘click’ chemistry. After full analytical and spectroscopic characterization of the ligands, their complex formation behavior will be investigated for nonradioactive or long-lived nuclides of the targeted metal ions and afterwards also for radiometals for PET and SPECT imaging like the ‘workhorse’ 99mTc.
Matteo Ippoliti
Quantitative characterization of tumor flow-metabolism mismatch by PET and MRI
PIs: Makowski, Brenner, Abram
Application area: Cancer
Modalities: PET, MRI, PET-MR
Background: Simultaneous acquisition of PET and MRI with a PET-MRI hybrid system has the potential to improve the accuracy of detection and quantitative characterization of flow-metabolism mismatch, as simultaneous acquisition avoids spatial registration errors and also eliminates additional variance associated with consecutive imaging at different time points. The project is centered on PET-MRI hybrid imaging for simultaneous acquisition of metabolism and blood flow/perfusion measures, as previous studies suggested that detection and quantitative characterization of flow-metabolism mismatch is clinically useful in defining the stage and potential responsiveness to treatment of various tumor entities.
Aim: To optimize the flow/metabolism acquisitions and gather several parametric maps (regional cerebral blood volume, regional cerebral blood flow and mean transit time) in order to quantitatively characterize flow/metabolism mismatch in tumors and identify biomarkers for treatment responsiveness.
Methods: The project is centered on PET/MRI hybrid imaging for simultaneous acquisition of metabolism (by the PET component) and blood flow measures (by the MRI component). Generation of the arterial input function for tracer kinetic modelling of (dynamic) PET in simultaneously acquired PET/MRI data. This will build on MRI-based delineation and motion tracking of a large artery within the acquisition field-of-view. Optimization of simultaneous FDG-PET/ perfusion weighted MRI for generation of parametric maps of regional cerebral blood volume, regional cerebral blood flow and mean transit time. Multivariate image analysis aimed at characterizing flow/metabolism mismatch.
Andreas Kofler
Deep Learning – Based Image Reconstruction for CT Quantification of Perfusion and Elasticity Related Parameters in the Heart
PIs: Dewey, Sack, Kachelrieß
Application area: Cardiovascular
Modality: CT
Background: Although CT accounted for only 9% of all radiological and nuclear medicine examinations in the year 2012, CT is responsible for around 62% of all medical radiation exposure in Germany (www.bfs.de). While temporally resolved CT techniques are very promising for improving our understanding of tissue perfusion such as that of the myocardium in coronary artery disease, current radiation dose levels to pursue dynamic CT are prohibitive for clinical use and require substantial reduction. In order to be able to perform a reliable quantitative estimation of perfusion parameters, one has to acquire high-dose images. Due to the high number of projections needed by the filtered back-projection method to deliver artifact-free images, one can, making specific assumptions on the underlying image-structure (e.g. sparsity), avoid to perform a high number of projections and instead apply iterative methods to reconstruct the images. Iterative methods usually work by alternating between reconstructing the images, also employing regularization techniques, and assuring their raw-data consistency. However, these methods tend to be time consuming due to the need of an iterative application of the forward and adjoint operator and have a limited dose-reduction potential.
Aim: To ensure a fast data acquisition procedure as well as a fast reconstruction of the images the work is going to focus on the applicability of Deep Learning methods for the task of the image reconstruction.
Methods: Deep Learning methods have nowadays become state-of-the-art methods for many tasks related to image analysis. Recently, Deep Learning methods have also been used for image denoising. In particular, in CT, artifacts resulting from undersampling show a specific structure which can be learned from a neural network in order to estimate artifact-free images. In the project, as a first step, we are going to train convolutional neural networks for the task of image denoising and evaluate their performance in terms of the predictability of perfusion parameters which are estimated from the artifact-free images.
Ledia Lilaj
Time-harmonic elastography for quantification of tissue pressure
PIs: Fischer, Sack, Hirsch/Jöhrens
Application area: MRI
Modalities: MRI and ultrasound
Background: Elastography is a biomedical imaging technique that visualises the differences in the biomechanical properties of normal and diseased tissues. In this field, several tissues have been analysed and some of them, like, for example, brain or liver tissues, are known to show poroelastic characteristics. Poroelastic models have been used to reconstruct tissue pressure-related parameters from time-harmonic displacement data as measured by Magnetic resonance elastography (MRE) in porous phantom materials.
Aim: The aim of this project is the development of elastography methods based on MRE or ultrasound which are sensitive to tissue pressure.
Methods: In a first step, I will develop an appropriate poroelastic phantom for MRE investigations. I will then develop an MRI sequence which allows the separate measurement of fluid and solid displacement data. By this sequence and the selected phantom I will analyse first the “effective medium” displacements, and second, the water and the solid displacements separately. After the phantom study, in vivo experiments will be done in order to assess the technique in the liver or brain imaging.
Jan Macdonald
Machine learning approaches for MR-fingerprinting of cardiovascular flow
PIs: Kutyniok, Schäffter, Schulz-Menger
Application area: Cardiovascular
Modality: MRI
Background: Recently, there has been an increased interest in employing machine learning methods (e.g. deep neural networks) for tasks in medical imaging such as image reconstruction, denoising, or super-resolution. Although first experimental works show promising results, there is yet an unsatisfactory lack of fundamental theory and understanding of these methods. A more rigorous mathematical analysis is needed before they could be considered admissible for clinical applications.
Aim: Development of a mathematical theory of deep learning for inverse problems. A particular focus lies on inverse problems arising in medical imaging, in particular the reconstruction of MRI signals from incomplete k-space data and MR parameter mapping in the framework of MR-Fingerprinting. As a first step, the basis pursuit problem (which arises in compressed sensing) is considered.
Methods: MR-flow is a well-established quantitative technique in diagnosis of cardiovascular medicine at all stages of life. However, due to the high spatiotemporal resolution requirements, fast and accurate flow quantification remains a challenge. Sometimes the differentiation of neighbouring arteries and veins can be difficult. Magnetic Resonance Fingerprinting (MRF) is a new approach that aims to acquire parameters of interest, such as T1, T2 and proton density, within one single scan. For this, unique signal evolutions (‘fingerprints’) of each tissue type are created and measured signals are matched to a signal dictionary to reconstruct parameters. For far, MRF has been applied to measure parameter maps of spin-density, off-resonance and relaxation times (T1, T2). Recently, MRF has also been investigated for perfusion parameter mapping. An important prerequisite for MRF is the dictionary, which is determined by simulating the signal evolution of the fingerprinting sequence setup (i.e., timing, flip angles etc.) using the Bloch equations. Matching of the measured data with the dictionary is usually performed voxel-vise based on least squares correlation. However, matching algorithms are time-consuming and prone to errors if parameter sampling of the dictionary is too sparse or encoding does not allow proper classification.
Johannes Mayer
Quantitative Assessment of Coronary Plaques by Motion-Compensated PET-MRI
PIs: Schäffter, Brenner, Makowski/Flöel
Application Area: Cardiovascular
Modality: PET-MRI
Background: Coronary plaques are the main cause of impaired tissue perfusion and subsequent myocardial infarction. Early detection of plaques likely to rupture could enable initiation of preventive treatment before stroke or infarction occurs. PET imaging using 18F-sodium fluoride (18F-NaF) has been shown to allow detection and assessment of coronary plaques. 18F-NaF accumulates in micro calcifications of plaques in the coronary arteries and identifies these high-risk plaques with high specificity. However, the diagnostic quality of 18F-NaF PET can be strongly impaired by physiological patient motion. Respiratory and cardiac motion lead to very large displacements of the coronary arteries compared to their diameter. Therefore, the PET signal of these small structures gets strongly blurred and cannot be distinguished from the background tissue anymore. PET-MR systems possess the ability to truly simultaneously acquire data of the two modalities. MRI offers a high spatial resolution in combination with an excellent soft tissue contrast which allows for extracting information on patient motion during the PET scan. This establishes the basis to use the complementary information provided by the MRI to improve the quality of the PET data.
Aim: Development of a more accurate and more reproducible quantification of risks posed by coronary plaques using PET-MR.
Methods: During this project, a novel high-resolution 3D MRI acquisition technique will be developed to obtain accurate cardiac and respiratory motion information. Image registration algorithms will be assessed to estimate the cardiac and respiratory motion of the heart. These will be incorporated into MR and PET image reconstruction using patient specific motion models. In particular, the project will focus on reconstruction methods making use of the entirety of the combined PET and MR data to mutually improve the image quality. The developed methods will be evaluated in simulations and studies with healthy volunteers and patients.
Azadeh Mohtashamdolatshahi
Towards quantitative structure-sensitive MPI: Application for sentinel lymph node detection
PIs: Schnorr, Taupitz, Schäffter, Käs
Application area: Cancer
Modalities: MPI, MRI
Background: The novel Magnetic Particle Imaging (MPI) modality detects and localizes magnetic nanoparticles (MNP) with high sensitivity. MPI has the potential to quantitatify MNP and their interaction with tissue environments, such as adhesion of MNP to macromolecular, ECM-, and cell surface components as well as internalization by phagocytosing cells. Experimental interstitial lymphography (IL) with sentinel lymph node targeting offers an ideal model to study interactions of MNP with different environments on a slow time scale and to study how these interactions influence the MPI signal.
Aim: Quantification of MNPs and their interactions with various environments in models of IL.
Methods: MNPs in various media and environments such as (in vitro models) water, blood, serum, lymph fluid, gel assays, cell cultures, (ex vivo models) lymph nodes and (in vivo models) injection site, lymph vessels, lymph nodes. Correlation with MPI, Magnetic Particle Spectroscopy (MPS), magnetorelaxometry, MRI, light scattering, iron analytics, histology, transmission electron microscopy (TEM).
Felix Schrank
Time-resolved quantification of myocardial microstructures by in-vivo viscoelasticity measurements
PIs: Sack, Schulz-Menger, Ittermann
Application Area: Cardiovascular
Modalities: MRI, Ultrasound
Background: Cardiac work is linked to the alteration of myocardial shear modulus. Current imaging markers of cardiac failure cannot measure forces and are limited in assessing the mechanical function of cardiac tissue. Cardiac elastography is sensitive to myocardial shear modulus and hence to mechanical relaxation abnormalities of myocardial tissue. However, current cardiac elastography does neither account for viscoelasticity nor anisotropy of myocardial tissue and is thus non-quantitative.
Aim: In the first period of my project I will focus on development and characterization of reference phantom materials which feature tissue-like viscoelastic properties.
Methods: The materials should be visible in MRI and ultrasound in order to compare the findings from ultrasound based time-harmonic elastography (THE) and multifrequency MR elastography (MMRE). The project will be continued with in-vivo cardiac MMRE and cardiac THE in healthy volunteers. Current MRE pulse sequences will be customized to the needs of cardiac elastography to enable a high spatially and temporally resolved quantification of myocardial shear modulus. Next, I will focus on anisotropic shear modulus recovery based on three-parameter inversions accounting for transverse isotropy, followed by a comparison of isotropic and anisotropic inversion and, tests of consistency and reproducibility of both. Finally, it is planned to compare the results of cardiac elastography with other imaging markers, such as volumetry, hemodynamic parameters obtained from 4D flow and myocardial perfusion.
Yavuz Oguz Uca
Molecular imaging of inflammation beyond macrophages by magnetic nanoparticle-enhanced MRI
PIs: Taupitz, Schroeder, Makowski
Application area: Cardiovascular
Modality: MRI
Background: A vast majority of diseases such as atherosclerosis or arthritis take place through inflammatory processes. Advances in the field of molecular imaging have enabled the use of nano-targeting concepts for detection and quantification of inflammation, thereby leading to better understand the critical cellular and molecular mechanisms behind diseases. In this scope, an important focus has been established on exploring the accumulation of phagocytic macrophages in the affected region by magnetic nanoparticle (MNP)-based magnetic resonance imaging (MRI). However, increasing understanding of the pathogenesis of such diseases suggest thorough investigation of major tissue structure alterations of the extracellular matrix (ECM) during the onset and progression of inflammation. Recent research has presented evidence on the accumulation of proteoglycan (PG) and glycosaminoglycan (GAG) molecules that are attractive sites for transchelation of electrostatically stabilized MNPs (ES-MNP) due to their negatively charged molecular structures as well as their ability to act as modulators to activate various information transmitting mechanisms. Such characteristics of these molecules make them excellent targets for molecular MRI. It has been shown that in vivo targeting of sugar-based components of ECM occur within minutes following the administration of MNPs while targeting of phagocytic cells takes hours to days. This could potentially aid clinical workflow and enhance the accuracy of imaging-based quantification methods, thereby offer valuable opportunities for biomedical research and clinical applications.
Aim and Methods: This project aims at: 1) Molecular MRI in small animal models for the detection of ECM changes in inflammation, 2) Monitoring local accumulation of suited imaging probes by in vivo and ex vivo examinations, 3) Assessment of mechanical properties of the inflamed tissue, and 4) correlation of imaging probe based and elastography based methods.
Stephan Wäldchen
Robust recovery strategies for the acoustic inverse scattering problem in anisotropic systems
PIs: Kutyniok, Fischer, Hege
Application areas: Cardiovascular, cancer
Modalities: PAT, US, MR
Background: The problem of inverse scattering appears in multiple ways in medical imaging. Photoacustic Imaging (PAT), ultrasound, and elastography all are all based on the scattering of acoustic waves in biological tissue. We want to backtrack the scattering patterns to the underlying tissue properties. This tissue is generally anisotropic and can include brain folds, blood vessels or lactiferous ducts. To increase the robustness to nois our reconstruction algorithms include regularization teqniques fit to this anisotropic geometry (e.g. shearlets).
Aim: At the moment we are focusing on modeling the 1-dim Helmholtz equation and inverting it for inhomogenous media using the stacked wave inversion. This means we are exciting waves with different frequencies and thus overdetermine the linear system for the stiffness of the tissue. Regularization is done with wavelets. Questions that arise here are sensibility to noise, frequency, distance of the frequencies and strength of the regularization. Once we move to 2-dim waves we will try regularization based on shearlets. After that we want to apply the methods to the real world data of MRE brain scans. It is not yet clear whether we want to go for high resolution or diagnostically useful stiffness values.Inverse scattering problems occur in medical imaging in various ways. Of particular importance is the acoustic inverse scattering problem, which models the reconstruction procedure from data generated by, for instance, PAT, US, and elastography.
Methods: We will introduce a mathematical model for 3D scatterers by accounting for anisotropic features prevalent in biological tissues such as connective tissue fibers, blood vessels, or lactiferous ducts. Given generalized geometrical features of self-similar 3D anisotropic networks in biological tissues, the model will provide an efficient mathematical treatment of highly complex scattering phenomena and enable us to invoke adapted regularization strategies such as the L1-norm of carefully chosen representation systems (e.g. shearlets) capable of sparsifying the model functions. Based on this approach, an optimal – in the sense of approximation accuracy – reconstruction scheme will be developed and analyzed with respect to realistic noise assumptions.
Patrick Werner
Quantification of alterations in ECM composition by switchable Xe-MRI reporters
PIs: Schröder, Taupitz, Bick
Application areas: Cancer, extracellular matrix, osteoarthritis
Modality: MRI
Background: Extracellular markers like matrix metalloproteinases (MMP´s) could provide important diagnostic information related to tumor growth, invasion and metastasis. However, relaxivity-based contrast agents do not provide enough sensitivity to visualize related processes like proteolytic degradation of ECM and alteration of cell-cell/ECM interactions.
Aim: It has been shown that the survival rate of patients increases dramatically with an early stage disease detection at the earliest possible time point. By achieving this goal we can focus on the preservation of health and not on the recovery.
Methods: Xe-biosensors have the potential to detect specific disease markers at nanomolar concentrations. The combination of Xe that underwent spin exchange optical pumping (SEOP) and the indirect detection by using Chemical Exchange Saturation Transfer (Hyper-CEST) enables a theoretical 10^7-fold signal enhancement compared to normal MRI measurements. A big advantage is that the contrast which is generated can be switched on and off through the MRI pulse sequence and is not masked by the background MRI signal. Based on the group expertise in designing Xe-biosensors and by using novel hyper-CEST methods we will pursue a two-fold approach to detect ECM changes: a) by addressing enzymes such as MMPs that are the cause of the degradation and b) by visualizing the loss of certain collagens that are result of ECM alteration. The first part will be based on novel Hyper-CEST methods using a smart sensor activated by MMP-induced hairpin cleavage. The second aspect of this project will be the detection of collagen II in the context of osteoarthritis and the interaction of MMP2, MMP9 with collagen IV due to related ECM digestion. This also links to the migration of metastatic cells in different animal cancer models. To support our results, the quantification of the ECM structural changes will be correlated to histology and biophysical methods.
